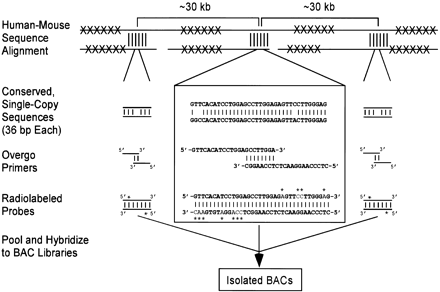
Strategy for designing universal overgo hybridization probes based on human-mouse sequence alignments. Orthologous human and mouse genomic sequences are masked for repetitive elements (indicated by Xs) and then aligned. Regions with high sequence conservation (indicated by vertical lines) are identified, and a single 36-bp human sequence from each is chosen based on GC content, percent human-mouse sequence identity, and uniqueness in the human genome (see Methods). A subset of these sequences then is chosen to optimize for interprobe spacing (∼30–40 kb). Three such conserved sequences are depicted in the figure, with greater details provided (in the box) for the middle one. Overlapping pairs of oligonucleotide primers are synthesized for each sequence and used to generate double-stranded, radiolabeled (indicated by *s) probes. The probes across a target region(s) then are pooled and used to screen arrayed bacterial artificial chromosome (BAC) libraries, allowing the isolation of individual positive BACs.











