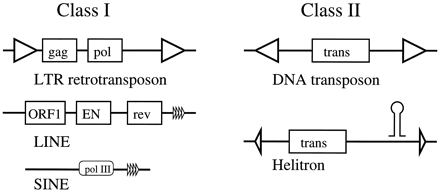
Some Known Types of Transposable Elements
Transposable elements are DNA sequences that move or are copied from one genomic location to another (Feschotte et al. 2002). They can be classified according to their transposition intermediate, RNA (class I) or DNA (class II), and whether they code for genes that catalyze transposition (autonomous TEs) or require these genes to be provided, usually by other TEs in the host (nonautonomous TEs). The genes required for autonomy are different for class I and class II transposons.
Class I TEs are transcribed to mRNA and then reverse-transcribed into a new locus. These include long terminal repeat (LTR) retrotransposons, close relatives of retroviruses with LTRs, requiring gag (capsid) and pol (protease, reverse transcriptase, integrase) genes for autonomy. The usual difference between LTR retrotransposons and viruses is the absence of an env gene, which allows viruses to breach host cell membranes and survive in the extracellular matrix. Having said this, some class I TEs are active retroviruses, such asDrosophila's gypsy (Mejlumian et al. 2002). Other kinds of class I TEs include long and short interspersed nuclear elements (LINEs and SINEs), the former of which containgag-like, endonuclease and reverse transcriptase genes, the latter a pol III promoter; both end with a short repeat.
Class II TEs, in contrast, are excised and reinserted as DNA. Characterized by terminal inverted repeats (TIRs), these elements can be autonomous with but a single transposase gene, which must specifically recognize the TIRs and catalyze the cut and paste transposition reaction (van Luenen et al. 1994).
A new type of eukaryotic TE called aHelitron, which was tentatively characterized as a class II (DNA) transposon but predicted to have a distinctive transpositional intermediate, has recently been discovered in the genomes of C. elegans, Arabidopsis, and rice (Kapitonov and Jurka 2001). Helitrons show sequence and structural homology to bacterial rolling-circle transposons, which transpose by a three-step mechanism: nuclease cut, strand transfer, and repair (Mendiola et al. 1994). They have short terminal repeats and specific sequences at each terminus (including a short palindromic signal at the 3′ end that may form a DNA stem–loop) direct the targeted cleavage reactions of transposition.
Nonautonomous TEs can be any DNA sequences containing transpositional-activating signals specifically recognized by proteins from autonomous TEs. Typically, nonautonomous elements are derived from autonomous elements by deletions or other mutations.
Nontransposable repetitive DNA includes local repeats and microsatellites caused by replicative errors such as polymerase stutter, as well as local duplications and larger satellite runs caused by chromosome mispairing during meiotic recombination.
The above diagram is not to scale. Exon structure and promoters are not shown (except the pol III promoter in SINEs). TEs may contain additional genes and regulatory sequences.











