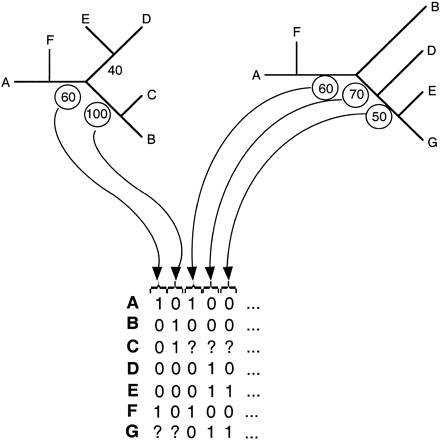
Construction of supertrees by MRP with bootstrap weighting. Each tree obtained for a set of species from a single orthologous gene family was coded into a binary matrix of informative sites. Only branches having a bootstrap value (or RELL-BP value for ML trees) over 50% were coded. The matrices obtained were concatenated into a supermatrix in which species absent from a gene family are encoded as unknown state (“?”). The supertree was calculated on the supermatrix usingDNAPARS with all default options, and 500 replicates of bootstrap were made using SEQBOOT.











