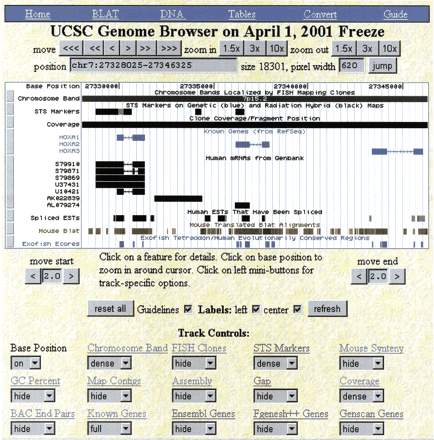
Part of the HOXA cluster as viewed in the University of California, Santa Cruz (UCSC) genome browser. The shortcut bar in blue provides quick access to BLAT searches, the DNA sequence, the annotations as text tables, earlier or later assemblies the genome, the corresponding NCBI and Ensembl views, and the user's guide. The controls directly beneath position the browser over a specific region in the genome. The large white picture in the middle displays various annotations. At the bottom are controls for fine-tuning the display and for the individual tracks. Only the first 15 of 31 available tracks are shown here.
This region contains three known genes that are all transcribed on the reverse strand as indicated by the arrowheads in the introns. Note the alternative splicing of HOXA1 in the Human RNA track. The Spliced EST track indicates that there is active transcription of a region between HOXA1 and HOXA2. Expressed sequence tag evidence for the presence of additional nonannotated genes in well studied regions like this often can be found using the UCSC browser. The Mouse Blat track indicated a high level of conservation between mouse and human in this region. Both the Mouse Blat and the Exofish ecores are based on translated alignments, but in highly conserved regions such as this it is not unusual for even translated alignments to paint conserved noncoding regions. The noncoding regions have diverged considerably more between human and pufferfish than between human and mouse.











