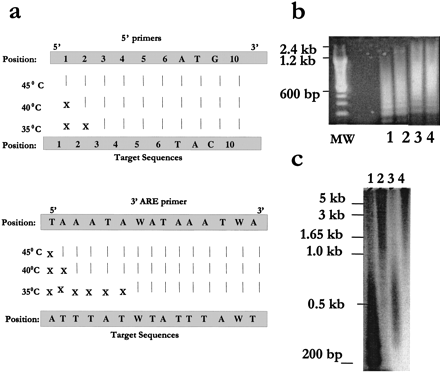
Characteristics of primer-target annealing and size distribution. (a) The annealing specificities of our primers under different annealing temperatures (35°C, 40°C, and 45°C) at the optimized ARE-cDNA PCR conditions (40 ng cDNA template, anti-Taq start conditions, 1 U Taq, 50 μM dNTP, and 30 cycles of amplification) were assessed. Twenty-six target sites for the 3′ and 5′ primers from 13 ARE-PCR products that were found to be portions of several known genes and genome/EST hits retrieved by BLASTsearch (Table 3) were evaluated. Vertical lines designate perfect matches (all at the 3′ end of the primer), and X designates mismatches. (b) Size distribution of amplified ARE-cDNA products. To visualize ethidium bromide smear for size distribution in agarose gels, amplified ARE-cDNA products (using an extension time of 3 min) were subject to a second ARE-cDNA PCR (Fig. 1) using 40μM dNTP. Samples were from THP-1 (lanes 1 and 3) and THP-1 treated with LPS + CHX (lanes 2 and 4) using two different 5′ primers (Ac and Gg; Table 1). MW: size markers. (c) Size distribution of the amplified radio labeled ARE-cDNA products. ARE-cDNA PCR from THP-1 samples, labeled with (α-32P)-dCTP, was performed at the optimized conditions (above) in the absence of (lanes1 and 3) or presence of 0.1 U of pfu (lanes2 and 4). Southern blotting was performed and autoradiogram was subsequently developed. Column purification was applied (lanes 3 and 4) to remove access dNTPs, primers, and short PCR products.











