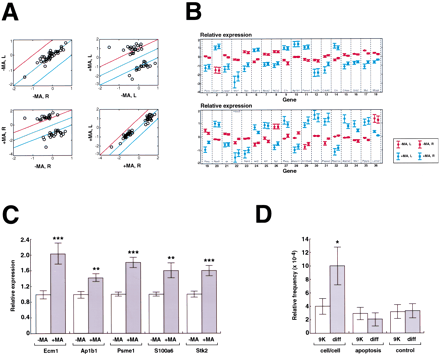
Differentially expressed genes. (A) Scatterplots show mean expression levels across the 40 voxels in the normal and MA brains on a logarithmic scale (log2) for genes wherep < 10−9. The comparisons within brains (left and right normal; left and right MA) provide another assessment of replicability (for normal brain, r = 0.90,F [1,34] = 143.98, p < 0.0001; for MA brain, r = 0.98, F [1,34] = 817.76,p < 0.0001). The comparison between brains (left normal and left MA, right normal and right MA) shows the differences in expression between genes repressed and induced in the MA brain. Red line: twofold above equivalent expression level; green line: equivalent expression level; blue line: twofold below equivalent expression level. (B) Mean expression levels across the 40 voxels of the normal and MA brains on a logarithmic scale (log2 ±SEM). (Red) Normal; (blue) MA. The genes are ranked from most (gene 1) to least significant (gene 36; p < 10−9). To allow replicability to be assessed, the data for the left and right halves of each brain were separated out as the left and right data points, respectively, for each gene. No significant differences were found comparing left normal with right normal (two-tailed t test on log2-transformed data, t = 0.012, df = 70,p = 0.99) and left MA with right MA (two-tailed ttest on log2-transformed data, t = −0.33, df = 70, p = 0.75). (C) Confirmation of gene expression differences for selected genes by real-time QRT-PCR analysis of voxel RNA. Expression levels are shown relative to controls and normalized using 18S RNA. The bars indicate mean ± SEM. (***)p < 0.001, (**) p < 0.01, two-tailedt-test. (D) Gene categories in the entire 9000-gene data set (9K) and in those genes significantly different (diff) between the control and MA brains (p < 0.001) when averaged across all 40 voxels. Genes involved in cell/cell interactions (cytoskeleton, extracellular matrix, and cell adhesion) were significantly overrepresented in the regulated subset (p = 0.02), but apoptosis-related genes were not (p = 0.39). The bars show mean ± SD, (*) p < 0.05, as judged using Monte Carlo statistics. The control category represents 15 randomly chosen genes to show the validity of the Monte Carlo analysis. As expected, this category showed no significant difference in frequency between the 9K and regulated data sets (p = 0.48).











