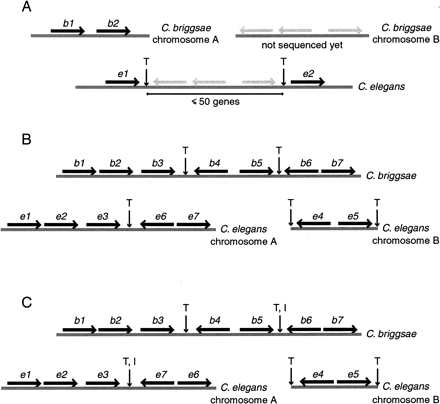
Method of detecting inversions and transpositions. (A) To detect transpositions to or from unsequenced parts of theCaenorhabditis briggsae genome, we looked along C. briggsae contigs for adjacent genes b1 and b2whose Caenorhabditis elegans orthologs e1 ande2 are on the same chromosome, where between e1 ande2 there are 1–50 C. elegans genes with unknownC. briggsae orthologs. We assumed that the genes between1 and 2 have transposed in either C. briggsae or C. elegans. (T) Transposition breakpoints. (B) To detect transpositions to or from sequenced parts of theC. briggsae genome, we looked along C. briggsaecontigs for three conserved segments in a row, where in C. elegans the first and third segments were close together on the same chromosome, and the middle segment was far away on the sameC. elegans chromosome or on a different C. eleganschromosome. We assumed that the middle segment (genes4–5) had transposed in either C. briggsaeor C. elegans. (C) To detect inversions, we looked along C. briggsae contigs for three conserved segments in a row, where in C. elegans the first and third segments were close together on the same chromosome, and the middle segment was far away on the same C. elegans chromosome or on a differentC. elegans chromosome, and either the first or third segment, or both, had inverted in either C. briggsae or C. elegans. Here the third segment (genes 6–7) has inverted.











