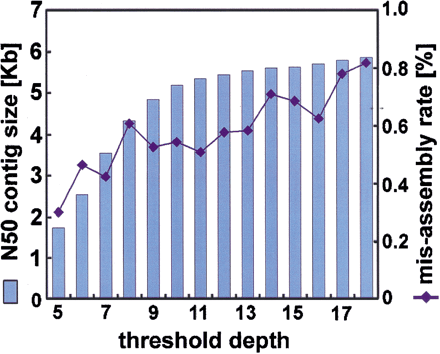
Tradeoff between contig size and accuracy of assembly. This analysis is based on the human 4× data set, using only repeat-maskedPhrap without the clone-end-pairing analysis. Increasing the threshold depth results in less of the sequence being masked, so the N50 contig sizes increase. For low-copy repeats, the resultant increase in misassembly rates is minor. The asymptotic contig size and misassembly rate, in the limit of no repeat masking, is somewhat larger than implied by this figure because transposon copy numbers run into the thousands, and this is well off the scale of the figure.











