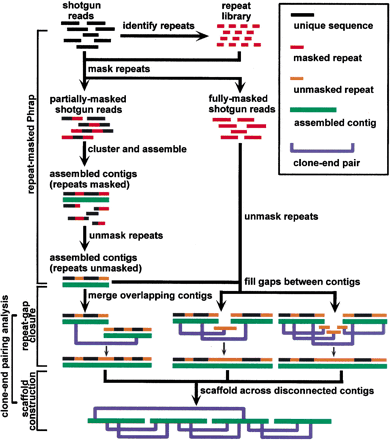
The RePS algorithm. Any 20mer that appears in the shotgun data set more often than a threshold depth is likely to be an exact repeat and is therefore masked out. Some sequence reads end up fully masked, but most have enough unique sequence in them to be used byPhrap. Repeat gaps are those for which the gap sequence is in the reads, but masked out by our procedure. LW gaps are those for which the gap sequence is not in the reads for statistical reasons (i.e., Lander-Waterman). Clone-end-pairing information is employed to help close the smaller repeat gaps. Large repeat gaps cannot be closed in this manner. Neither can LW gaps. But as long as the clone-insert sizes are larger than the remaining gaps, there is a reasonable probability that we can build scaffolds to bridge over the gaps and order and orient the contigs.











