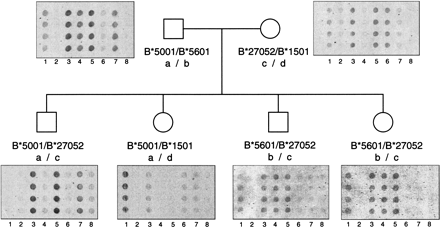
Segregation of HLA-B SNPs in a family study. Probes descriptive for positions 154–173 (lanes 1–4) and positions 194–213 (lanes 5–8) of exon 2 are illustrated for HLA-B*5001 (paternal “a” haplotype, lanes 3 and7), HLA-B*5601 (paternal “b” haplotype, lanes 4and 5), B*27052 (maternal “c” haplotype, lanes 3and 5) and B*1501 (maternal “d” haplotype, lanes1 and 6). Positions of the probes are numbered according to the HLA/IMGT database. Cross-hybridization is observed for eight of the 137 (6%) probes, however, these spurious signals can be identified by analyzing the hybridization results of the other probes in the group. For example, the two family members with the “b/c” haplotype in this figure demonstrate a false-positive hybridization with the probe in lane 1. Despite the signal from this probe, the correct allele was called because of the lack of signal from the probe in lane 6.











