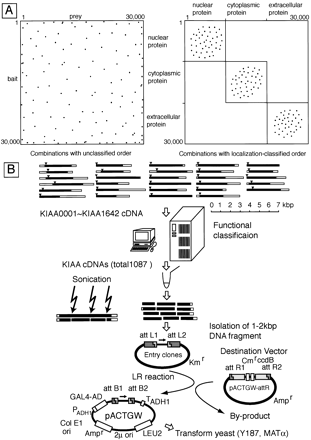
Diagram of our strategy for constructing a functionally classified library composed of long cDNAs. (A) Illustration of different combinations according to functionally classified order. Because there are ∼30,000 human proteins, our ultimate aim would be to prepare a complete protein–protein interaction map by assessing interactions between ∼30,000 × ∼30,000 protein combinations. One dot represents the interaction between one vertical-ordered protein (i.e.,left, ordinate) and one horizontal-ordered protein (i.e.,left, abscissa). The set of possible interactions to be assayed is extremely large. If proteins are, instead, first categorized according to their location and function, as shown in theright panel, interaction dots are clustered to smaller regions, and thus the set of possible interactions to be investigated is reduced considerably. (B) Schematic diagram of library construction. Black and white bars represent open reading and untranslated regions, respectively. Triangle indicates the first methionine codon. After functional classification in silico, randomly fragmented cDNAs were introduced into vectors for yeast two-hybrid screening using the Gateway system.











