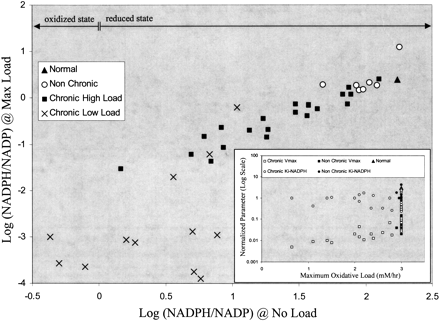
The ability of glucose-6-phosphate dehydrogenase (G6PD) variants to respond to oxidative load based on in silico red blood cell model results. The NADPH/NADP is plotted for maximum load (vox = max value tolerated) versus no oxidative load (vox = 0) for each G6PD variant. The nonchronic cases cluster near the normal case and can sustain a maximum oxidative load of near normal (3 mM/h). The chronic cases can be split into two categories, those that can sustain a near-normal load (chronic high load) and those who are well below normal (chronic low load). The horizontal arrows demarcate two regions: an oxidized state (Log [NADPH/NADP] < 0) where the majority of the cofactor is in the form of NADP and a reduced state (Log [NADPH/NADP] > 0) where the majority of the cofactor is in the form of NADPH. For G6PD enzymopathies, the oxidized state is prevalent, while the reduced state is found in normal individuals (Kirkman et al. 1975). The insert shows no clear correlation between Vmax, Ki-NADPH, and the cell's ability to withstand oxidative loads.











