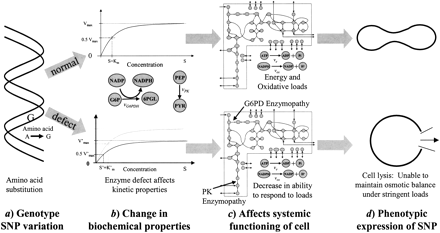
A schematic depiction of the genotype-phenotype relationship; from sequence variation (a) to changes in biochemical function (b) to network properties (c) to overall physiological function (d). The upper path signifies a normal pathological situation. The DNA sequence codes a nondeficient enzyme that catalyzes the glucose-6-phosphate dehydrogenase or pyruvate kinase reactions normally resulting in homeostatic red blood cell metabolism with the cell assuming its traditional biconcave shape. The lower path represents a pathological situation, a defect in the DNA sequence that leads to a metabolic enzyme with altered catalytic activity. The result is a loss in the cell's ability to respond to oxidative and/or energy loads, hence an inability to maintain osmotic balance and stability that ultimately results in lysis–the phenotypic expression of the genetic defect.











