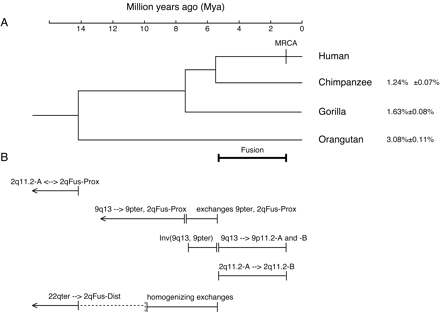
Estimated timing of duplications, inversions, and relocations of 2qFus-paralogous blocks based on sequence identity measures and fluorescence in situ hybridization analyses of hominids. (A) Phylogeny of hominids. The branch lengths and estimated speciation dates are based on sequence analyses of 53 autosomal intergenic nonrepetitive DNA segments analyzed by Chen and Li (2001). The speciation dates are drawn at the midpoint of estimated ranges: human–chimpanzee, 4.6–6.2 Mya; human–gorilla, 6.2–8.4 Mya; human–orangutan, 12–16 Mya. MRCA marks the estimate of the time of the most recent commonancestor of all modern humans (Yu et al. 2001). The average ± SD percent sequence substitution (Jukes-Cantor model, excluding indels) between human and each of the three other hominids is given at the right (Chen and Li 2001). (B) Estimated timing of events involving 2qFus-paralogous blocks. The blocks are identified by their current positions in humans (i.e., 2qFus-Dist and 2qFus-Prox are the regions forming the distal and proximal sides of the 2q fusion site, respectively; both were at chromosome tips when the duplicative transfers occurred). Ancestral and derived states, when indicated, are inferred from breaks that disrupt genes, specific repetitive elements, or isochore patterns; the copy carrying the full-length gene/element and/or lacking an isochore transition at the breakpoint is assumed to represent the ancestral version.











