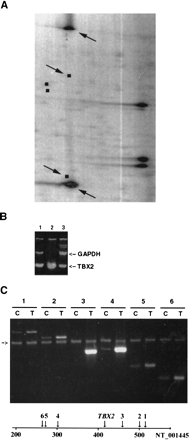
(A) Tumor 200 NotI/EcoRV/HinfI restriction landmark genomic scanning (RLGS) profile. Black squares represent predicted spots from chromosome 17 in this section of the RLGS pattern. Arrows point to two amplified fragments specific to Tumor 200 and to the location of their predicted fragments from chromosome 17. The three highly intense spots in the right part of the gel represent rDNA. (B) PCR analysis of TBX2 genomic amplification. Lane 1: MCF7; lane 2, Tumor 200; lane3, Control DNA. Arrows point to GAPDH (407 bp) andTBX2 (188 bp) PCR products. (C) Extent of the 17q23 amplified region as determined by PCR analysis. Lanes 1– 6represent multiplexed PCR using GAPDH and primers 1–6, respectively (C, control DNA; T, Tumor 200 DNA). An arrow points to the location of GAPDH (407 bp) PCR product. Positions of primer pairs 1–6 and TBX2 along sequence NT_001445 are drawn. The scale is given in kb.











