Optimization of Neural Network Architecture (Input Parameters vs. Performance on the EGF-Like Domain Type)
| Training set | Test set | Total | ||||||||||||||||||||||||||||||||
| Parameter | No | tp | fp | fn | tn | C | tp | fp | fn | tn | C | tp | fp | fn | tn | C | ||||||||||||||||||
| 1 | NSD, AVS | 2 | 242 | 23 | 49 | 311 | 0.77 | 125 | 3 | 20 | 137 | 0.85 | 360 | 14 | 76 | 460 | 0.81 | |||||||||||||||||
| 2 | NSD, AVS, P
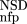 (NSD), P
(NSD), P
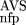 (AVS)
(AVS)
|
4 | 284 | 4 | 7 | 330 | 0.97 | 142 | 3 | 3 | 137 | 0.96 | 426 | 8 | 10 | 466 | 0.96 | |||||||||||||||||
| 3 | NSD, AVS, P
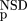 (NSD),P
(NSD),P
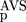 (AVS)
(AVS)
|
2 | 291 | 2 | 0 | 332 | 0.99 | 145 | 3 | 0 | 137 | 0.98 | 434 | 4 | 2 | 470 | 0.99 | |||||||||||||||||
| 4 | NSD, AVS, P
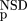 (NSD), P
(NSD), P
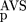 (AVS), P
(AVS), P
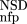 (NSD), P
(NSD), P
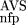 (AVS)
(AVS)
|
4 | 284 | 4 | 7 | 330 | 0.97 | 142 | 3 | 3 | 137 | 0.96 | 426 | 8 | 10 | 466 | 0.96 | |||||||||||||||||
-
↵tp, True positives; fp, false positives; tn, true negatives; fn, false negatives.
-
C is the Matthews (Pearson) correlation coefficient (Matthews 1975),
 Each neural network contained one hidden layer with the same number of elements as the number of input parameters. Note that
the Total values were obtained by retraining the ANNs on the entire dataset, so these values are not necessarily equal to
the sum of the corresponding Training Set and Test Set values.
Each neural network contained one hidden layer with the same number of elements as the number of input parameters. Note that
the Total values were obtained by retraining the ANNs on the entire dataset, so these values are not necessarily equal to
the sum of the corresponding Training Set and Test Set values.











