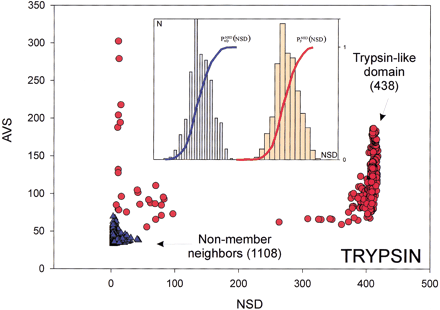
Variables used to characterize domain similarities. AverageBLAST similarity score (AVS) versus number of significant BLAST similarities (NSD) plot for Trypsin domains (red circles) from SBASE 7.0 (Murvai et al. 2000a). Each red circle represents a domain sequence.
The AVS andNSD values are computed from the BLASTsimilarities between the sequence and other members of the group. Blue triangles represent domain sequences that are not Trypsin
domains but still produce a significant BLAST similarity score with some of the Trypsin domains. The database versus database comparison was carried out with BLAST (Altschul et al. 1990), by use of a significance threshold of 0.8 and a minimum score of 32. (Inset) Schematic representation of the empirical distributions (see text for details). Values of P
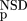 (NSD), and P
(NSD), and P
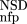 (NSD), are simply read out from the precomputed empirical distributions. A similar procedure is followed for the AVS scores. The resulting six values are the input parameters of the ANNs.
(NSD), are simply read out from the precomputed empirical distributions. A similar procedure is followed for the AVS scores. The resulting six values are the input parameters of the ANNs.











