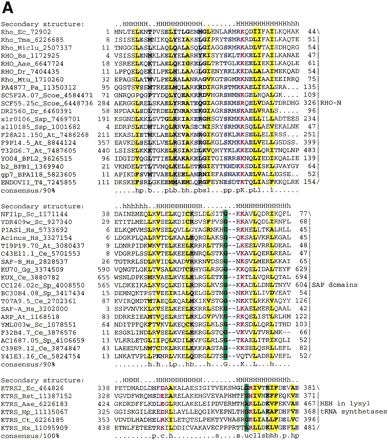
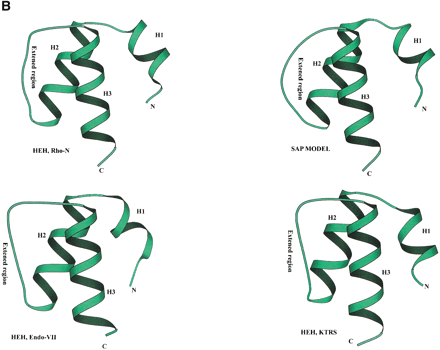
(A) Multiple sequence alignment of different classes of HEH domains. Each of the alignments is colored according to a separate consensus using the rules described in the legend to Figure 2. The secondary structure shown above the alignment was derived from the structures of Rho, Endo-VII and K-TRS. For the SAP domains, the structure was predicted using the PHD program. The species abbreviations are the same as in Figure 2; those not present in Figure2 are : Ec, Escherichia coli; Ssp, Synechocystis sp.; Tma, Thermotoga maritima; Dr, Deinococcus radiodurans; Aae, Aquifex aeolicus; BPL2, lactococcal Bacteriophage L2; BPA118, Listeria bacteriophage A118; T4, Bacteriophage T4; Miclu, Micrococcus luteus; Ce,Caenorhabditis elegans; Ct, Chlamydia trachomatis; Hp, Helicobacter pylori; Bst, Bacillus stearothermophilus. (B) Structures and models of different forms of the HEH domain shown in the alignment. The NH2 (N) and COOH (C) termini of the HEH domains are indicated.











