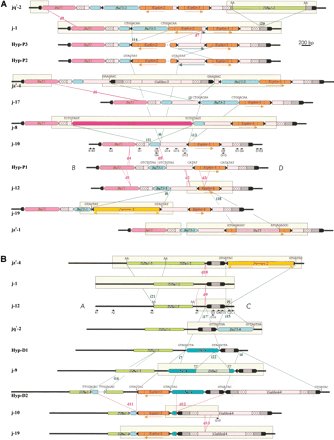
Schematic representation of the structures found at the proximal (A) and distal (B) breakpoints of inversion 2jin the 30 2j lines studied. All different structures are shown, except for that of j-16 in the proximal breakpoint, which differs from jz3–4 by the absence of d6 deletion. Thick lines represent the single-copy A, B, C, andD sequences. TEs are represented as colored boxes and sharp ends correspond to the ITRs. Insertions and deletions are delimited by green and red lines, respectively, and are named with an i or a d followed by a number. Target site duplications flanking the insertions are shown above them. Blue lines indicate the inversion of an internal segment. Arrows below the diagrams inform on the orientation of some homologous segments. Segments sequenced in each structure are enclosed within clear rectangles. Only the D. buzzatii lines representative of each structural variant are shown. Lines sharing the same structure in the proximal breakpoint are jq7–1, jq7–2, and jq7–3; j-1, j-2, j-3, j-4, j-5, j-6, j-7, j-14, j-15, j-20, j-21, and jq7–4; j-9, j-11, j-12, j-13, j-18, and j-22 (deletion d2 was detected during j-12 sequencing and we do not know whether it is present in other lines or not); jz3–1, jz3–2, and jz3–3. Lines sharing the same structure in the distal breakpoint are j-1, j-2, j-3, j-4, j-5, j-6, j-7, j-13, j-15, j-20, j-21, jz3–2, jq7–1, and jq7–3; j-8, j-11, j-12, j-14, j-16, j-17, j-18, j-22, jz3–1, jz3–3, and jq7–4. Hyp are hypothetical structures not found in our sample of 2j lines. Small black arrows are PCR primers used in the study.











