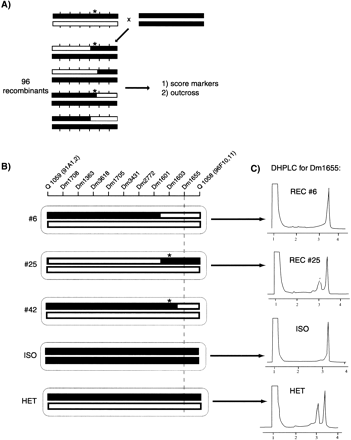
Recombination mapping of a recessive lethal mutation by using SNP markers. Chromosomes (bars) and molecular markers (vertical hatch marks) are shown. The mapping process occurs in two stages. (A) The mutation (asterisk) induced in the w;iso2;iso3 background (black bar) that has been mapped previously to a chromosome and balanced is mapped relative to a polymorphic mapping strain (open bar). Single flies heterozygous for the mutation-carrying chromosome and the mapping chromosome are crossed to flies homozygous for the parental w;iso2;iso3 strain to generate 96 recombinant flies. The four recombinant classes are represented. Each recombinant fly strain is assayed for a low-density set of markers that span the chromosome. These markers can be of any type, including SNPs or P-element insertions. We have typically tested six markers spaced at ∼10-Mb intervals on these 96 recombinants, for a total of 576 assays. Each recombinant also is assayed for presence or absence of the mutation by outcrossing. From the outcross data and initial marker data on this set of 96 recombinants, the mutation can be assigned to an interval of 10–20 Mb. (B) A higher density set of SNPs then is assayed on recombinants from A that break in the appropriate interval. An example of mapping a mutation in the hedgehog gene is shown. SNP markers are indicated by the STSs from which they are derived. Two P elements (Q 1059 and Q 1058) that were used to localize these mutations also are indicated. The chromosomal compositions of three recombinant (#6, #25, and #42) and two control (ISO and HET) flies are represented. Asterisks indicate chromosomes that carry the mutation as defined by outcrossing. The SNP markers shown were scored using DHPLC. The recombinants shown delimit the position of these mutations to between Dm1601 and Dm1655, a region of ∼984 kb. Subsequent complementation testing showed that these mutations are alleles of thehedgehog gene, which lies between Dm1601 and Dm1655. (C) DHPLC scoring of SNPs. PCR products were amplified from recombinants and analyzed under partially denaturing conditions. Data are shown for Dm1655, a C/T dimorphism. Run time in minutes is shown on the X axis and ultraviolet absorbance on the Y axis. Dm1655 was analyzed from the following strains: w;iso2;iso3 (ISO), a w;iso2;iso3/Q1059 heterozygote (HET), and hedgehogrecombinants 6 and 25 (#6 and #25). In this example, the sample from recombinant 25 shows a heteroduplex pattern and therefore is scored as a heterozygote. (DHPLC) denaturing high performance liquid chromatography.











