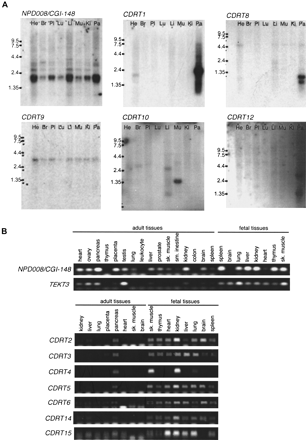
Expression studies of genes identified in the CMT1A/HNPP region. (A) Multiple-tissue Northern blot analyses. Tissues are indicated at top of each lane (He) Heart; (Br) brain; (Pl) placenta; (Lu) lung; (Li) liver; (Mu) skeletal muscle; (Ki) kidney; (Pa) pancreas. Marker sizes are 9.5, 7.5, 4.4, 2.4, and 1.35 kb. (B) Multiple-tissue RT–PCR analyses. NP008/CGI-148 is expressed in a wide variety of tissues. A 2-kb major transcript and two minor transcripts are observed. TEKT3 revealed no expression by Northern blotting (data not show), but high expression in testis by RT–PCR. Faint expression is also observed in both ovary and pancreas by RT–PCR. Fetal tissues reveal low level but ubiquitous expression.CDRT1 shows a major 2-kb and a minor 1-kb transcripts in pancreas. Faint expression is also identified in the heart.CDRT8 reveals two short transcript (0.7 and 1 kb) in the pancreas. A 3-kb transcript of CDRT9 represents low-level ubiquitous expression. CDRT10 is expressed in the skeletal muscle as a 1.3-kb transcript. CDRT12 reveals a 2.8-kb transcript in the pancreas at very low expression levels. CDRT2, CDRT3, CDRT4, CDRT5, CDRT6, CDRT14, and CDRT15 reveal no expression in adult tissues by Northern blotting (data not shown) and RT–PCR analyses, but show obvious expression in fetal tissues.











