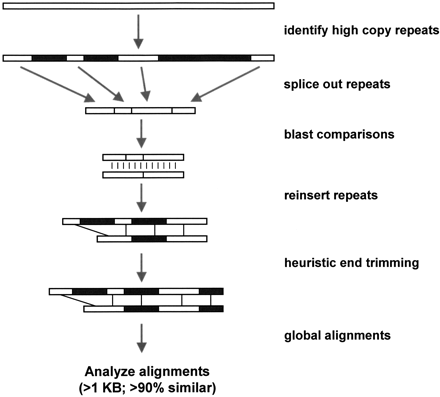
Detection Method. The method combines DNA sequence analysis software and a suite of Perl scripts that are optimized for the detection of large highly similar duplications. Briefly, the genome assembly (2.6 Gb) is broken into tractable 400-kb segments. For each segment, common repeats (blue) are identified with RepeatMasker. Repetitive sequence is then removed (“fuguized”) leaving putatively unique DNA. All fuguized pieces are then compared byBLAST. Repeats internal to an individual 400-kb segments are detected with BLASTZ. Relaxed affine gap parameters are used allowing gaps up to 1 kb in size to be traversed. Fuguized pairwise alignments (>0.87 similarity and >500 aligned bp) have their common repeats reinserted and then the alignment ends undergo heuristic trimming allowing for refinement of alignment end points which may lie within common repetitive sequence. The program ALIGNgenerates optimal global alignments from which final alignment statistics are calculated. Global alignments >1000 bases aligned and >90% identity were selected in this analysis.











