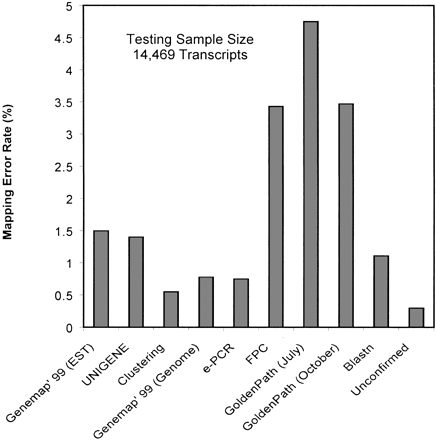
Estimated mapping error rates by the integration of multiple mapping data sets. A total of 14,469 transcript consensus sequences all containing the following mapping information were tested: the Genemap'99 for the ESTs contained in each of the clusters; the original UNIGENE cytogenetic or chromosomal assignments; the Genemap'99, fingerprint, and GoldenPath contigs (July and October versions), and the e-PCR maps for the genomic clones contained within the transcript consensus. Mapping errors were scored by an inconsistent chromosomal assignment of one map compared with five other maps. Nonspecific BLAST alignments were scored when the transcripts and the genomic clones putatively harboring the transcripts could be consistently mapped to different chromosomes. Errors occurring during the original UNIGENE clustering were detected when two distinct and consistent RH mapping assignments were found within the same UNIGENE clusters. Unconfirmed mapping errors were scored when more than one inconsistent mapping assignment was found for a given transcript.











