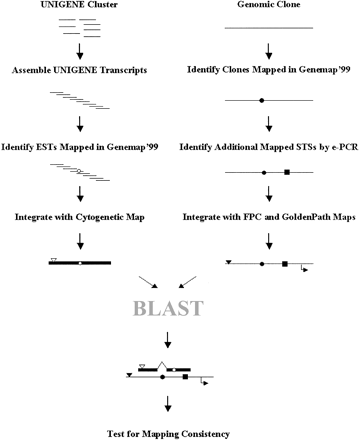
The data flow in mapping human transcript consensus sequences. UNIGENE clusters were assembled into transcript consensus sequences. The RH mapping information for the individual ESTs and other cDNAs available from Genemap'99 was incorporated into the transcript consensus sequences, along with the cytogenetic and/or chromosomal assignments available in the original UNIGENE data set. Mapping information for each genomic clone was compiled on the basis of the integration of four largely independent maps: Genemap'99, e-PCR and fingerprint maps, and GoldenPath sequence contigs. The resulting transcript consensus wasBLASTed against the mapped genomic clones. The final transcript map was substantiated by the integration of multiple maps available for both the transcripts and the genomic clones. (Open circles) Genemap'99 EST RH assignments; (open inverted triangles) UNIGENE cytogenetic or chromosomal assignments; (closed circles) Genemap'99 RH assignments for genomic clones; (closed boxes) additional mapped STSs; (closed inverted triangles) fingerprint-mapping assignments; (bent arrows) GoldenPath positional assignments.











