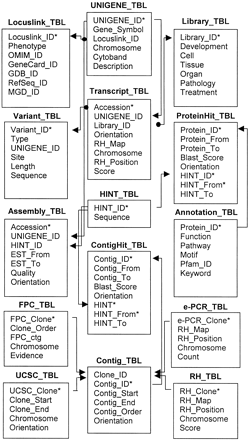
The database schema for the integration and mining of functional and mapping information of human transcript consensus. The UNIGENE_TBL and Locuslink_TBL were derived from the original UNIGENE and Locuslink data sets, respectively. The Transcript_TBL was created by the integration of UNIGENE and Genemap'99 databases. The cDNA library information in the Library_TBL was standardized further into appropriate categories: Development (e.g., embryonic stages), Cell, Tissue, Organ, Pathology (e.g., tumor), and Treatment. The Assembly_TBL defined the start and end positions and the orientation for each transcript in its contig, and its Smith–Waterman score compared with the consensus. All consensus sequences were stored in the HINT_TBL. The potential splicing variants were represented in the Variant _TBL by the site and length of the insertion or deletion for a given variant. The ProteinHit_TBL and ContigHit_TBL were generated by BLASTing the transcript consensus with the protein (SWISSPROT, PIR, and TrEMBL), and genomic (Ensembl) sequence databases, respectively. A majority of the functional annotations in the Annotation_TBL were derived from the SWISSPROT data set, supplemented with the key words from the PIR and TrEMBL protein entries. The Contig_TBL, based on the Ensembl database, was used to order and position individual sequencing contigs (see Methods). Mapping information for individual clones was integrated by use of four different maps: the assembled genomic contigs (UCSC_TBL); the fingerprint map (FPC_TBL); and the radiation hybrid maps (RH_TBL and e-PCR_TBL). (*) Primary or joint keys for each table; (arrows) one to many relationships; (arrows with closedcircles) one to one relationships.











