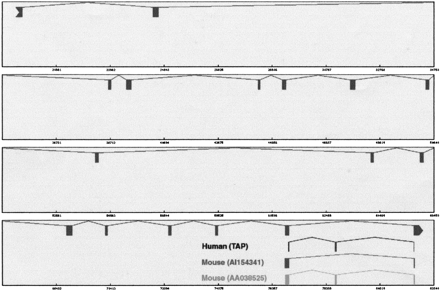
An example of conserved alternative splicing pattern. Shown here is a conserved exon skipping event at the 3′ end of RPN2 gene (NM_002951). The reference gene structure is displayed in the top level. The alternative splice pairs predicted from human ESTs, 76969–78662 and 78709–81563, are shown in the second level. We found that both the reference and alternative patterns were conserved. The mouse ESTs were aligned to the human genomic template using sim4. Each aligned block has ± 88% identity. The graphic plot was modified to illustrate these alignments. In the third level, the alignment of ESTAI154341 shows the same pattern as the reference gene structure. In the bottom level, the alignment of EST AA038525 displays the alternative splicing pattern.











