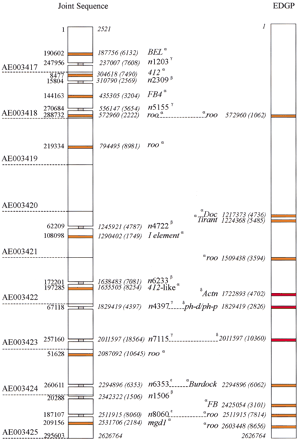
Sequence comparisons. A comparison of EDGP sequence of the tip of theX chromosome with that of the Drosophila Joint Sequence in the same region. The comparison was made using theMUMmer program (see Methods). The GenBank accession numbers corresponding to the Joint Sequence are shown on the left (AE003417–AE003425); note that this is part of a unitig (Myers et al. 2000). The blocks indicate regions of ⋝1 Kb present in one sequence but not the other. The position and length of each block of sequence ⋝1 Kb that is unique to one sequence is shown; each GenBank accession is numbered to the left of the unitig, the corresponding base position within the EDGP sequence is shown in italics to theright of the unitig. The EDGP sequence is numbered continuously. The length of each block of unique sequence is in parentheses. The nature of these sequence segments is shown in thecenter (note that a segment may include sequences in addition to those identified here). The segments corresponding to transposable elements are indicated in orange; those corresponding to known genes are red; a gray “neck” depicts a sequence interrupted by a large block of n's of length N (nN). The Greek superscripts (α,β,γ,ɛ,δ) refer to the class of sequence difference (see text). Note that there are an additional nine transposable elements in the EDGP sequence that are not seen to differ in the Joint Sequence.











