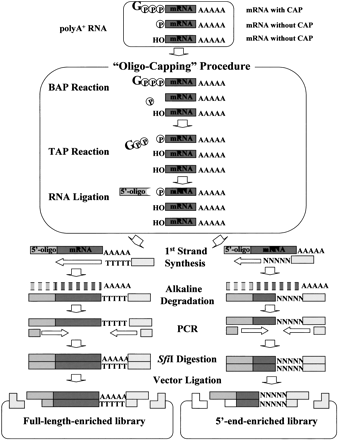
Schematic representation of the construction of oligo-capped cDNA libraries. The cap structure of the mRNA was replaced with the 5′ oligonucleotide by the oligo-capping method, which consists of three enzymatic reaction steps. Bacterial alkaline phosphatase (BAP) hydrolyzes the phosphate of the 5′ ends of truncated mRNAs whose cap structures have been degraded. Tobacco acid pyrophosphatase (TAP) removes the cap structure, leaving a phosphate at the 5′ end. T4 RNA ligase, which requires a phosphate at the 5′ end as its substrate, selectively ligates the 5′ oligonucleotide to the 5′ end that originally had the cap structure. Using oligo-capped mRNA, first-strand cDNA was synthesized with dT adapter primer. After alkaline degradation of the RNA, first-strand cDNA was amplified by PCR, digested with restriction enzyme SfiI, and cloned into a plasmid vector. For further details of the procedure, see references (Suzuki et al. 1997,2000). RNA and DNA molecules are represented by dark gray bars, the 5′ oligonucleotide by light gray boxes, and PCR primers by broken bars. (Gppp) Cap structure, (p) phosphate, (OH) hydroxyl.











