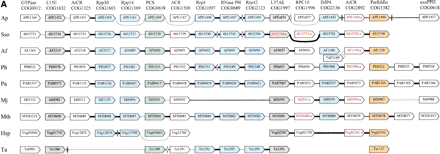

Organization of genes encoding predicted exosome subunits and functionally related proteins in archaeal genomes. (A) The potential exosomal superoperon. (B) Additional predicted operons coding for proteins functionally linked to the predicted exosome and the proteasome. Genes are not drawn to scale; the direction of transcription is indicated by arrows. The multiple gene-by-gene alignment was produced by manually combining template-anchored genome alignments; orthologous genes are aligned. For each column of the alignment, the number of the respective COG and the systematic subunit name or a functional designation are shown. Adjacent genes are connected with lines; thick lines indicate intergenic regions <20 nucleotides, thin lines those in the range of 20–50 nucleotides, and dotted lines those >50 nucleotides. The unconnected genes are located elsewhere in the genomes (which is also clear from the indicated gene numbers). The color coding shows functionally related groups of proteins: blue predicted exosome subunits (including the RNase P subunits Rpp30 and Rpp14), with blue hatching indicating tentative predictions (see text); green, proteasome subunits; gray, ribosomal proteins; gold, cotranslational chaperones; white, uncharacterized proteins and other functions, including flanking genes with no predicted functional connection with the exosome. The gene names shown in red and with the suffix a indicate predicted genes that are missing in the original genome annotation, but were identified during this analysis using TBLASTN searches. Diamonds show genes present in the original annotation that are inserted between the conserved genes; the open diamonds show predicted genes that significantly overlap with the conserved ones and are probably spurious; red diamonds indicate nonoverlapping genes that are likely to be real. Abbreviations: ACR, ancient conserved region; ArCR, archaeal conserved region; MTR, methyltransferase; PCS, proteasome catalytic subunit; PRS, proteasome regulatory subunit; exoPPH, exopolyphosphatase. The species abbreviations are as in Fig. 1. Hal,Halobacterium sp.











