
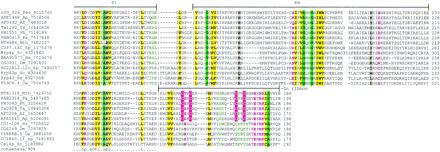
Multiple alignment of the Rrp4p and Csl4p subunits of the eukaryotic and predicted archaeal exosomes. The proteins are denoted by the gene names, Gene Identification (GI) numbers, and abbreviated species names. The positions of the first and the last residue of the aligned region are indicated for each sequence; variable spacers between the aligned blocks that were omitted from some of the sequences are indicated by numbers. The boundaries of the two predicted RNA-binding domains, S1 and KH, and the novel, amino-terminal pre-S1 domain are shown. The alignment coloring is based on the 90% consensus, which is shown underneath the alignment; b indicates a big residue (E,K,R,I,L,M,F,Y,W), h indicates hydrophobic residues (A,C,F,I,L,M,V,W,Y), a indicates aromatic residues (F,Y,W), s indicates small residues (A,C,S,T,D,N,V,G,P), u indicates tiny residues (G,A,S), p indicates polar residues (D,E,H,K,N,Q,R,S,T), and c indicates charged residues (K,R,D,E,H). The conserved cysteines that form a Zn-ribbon in the archaeal but not in the eukaryotic proteins are shown by white letters against a red background. The secondary structure elements predicted for the pre-S1 domain using the PHD program and a preconstructed multiple alignment as the input are shown above the alignment. H(h) indicates α-helix and E(e) indicates extended conformation (β-strand); upper case indicates the subset of the predictions with an estimated 80% confidence level. The species abbreviations are: Af, Archaeoglobus fulgidus; Ap,Aeropyrum pernix; Ce, Caenorhabditis elegans; Hs,Homo sapiens; Dm, Drosophila melanogaster; Mth,Methanobacterium thermoautotrophicum; Pa, Pyrococcus abyssii; Ph, Pyrococcus horikoshii; Ta, Thermoplasma acidophilum; Sc, Saccharomyces cerevisiae; Sp,Schizosaccharomyces pombe; Sso, Sulfolobus solfataricus.











