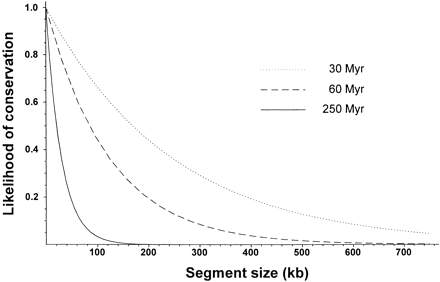
Potential transferability of positional information from Drosophila melanogaster to taxa at different phylogenetic distances. The probability that a chromosomal stretch with a relative length l has not been disrupted after the fixation of2n breakpoints isP = e −2nl. 2ndepends on the evolutionary divergence time between lineages compared. Divergence times indicated in the chart ×2 were taken from the comparisons among groups of species within the Sophophorasubgenus (Throckmorton 1975), among subgenera within theDrosophila genus (Spicer 1988), and among some of the main insect orders (Friedrich and Tautz 1997), respectively. A rate of 1.85 disruptions per Myr was assumed in all cases.











