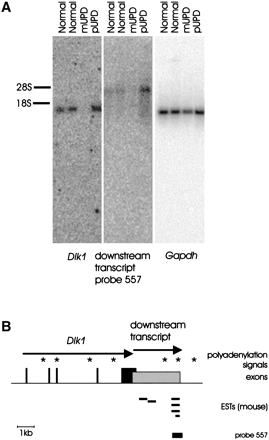
Imprinted expression and physical structure of a transcript downstream of Dlk1 in mouse. (A) Northern blot hybridization of enriched poly A+ RNA hybridized with probes specific for Dlk1, the transcript downstream of Dlk1, and Gapdh, respectively. The probe specific for Dlk1 spanned exons 1 and 2, the expression of the Dlk1 downstream transcript was tested using a genomic HpaII fragment overlapping with the 3– end of the transcript. Hybridization of Gapdh transcripts proved similar amounts of RNAs in all lanes. The positions of the 18S and 28S rRNA bands are indicated. (B) Dlk1 exons are shown as black bars. The Dlk1 downstream transcript overlaps with the last exon of Dlk and is shown as a grey box, starting at the most upstream 5′ end predicted by the 5′RACE experiment. Mouse ESTs that indicated the presence of a transcript downstream ofDlk1 are shown as horizontal bars. Asterisks represent polyadenylation signals. The position of the HpaII fragment used as a probe for the detection of transcripts downstream ofDlk1 is indicated. The schematic is drawn to scale.











