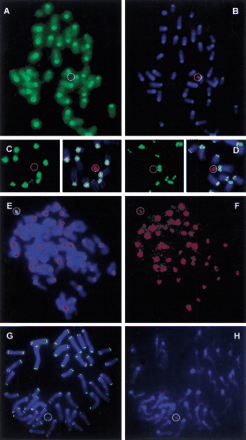
Characterization of the HCV in F1 transchromosomal mice. Metaphase spreads of tail fibroblasts of HCV+ F1 mice were used for FISH. A cohybridization was performed with a biotinylated mouse Cot1 DNA probe (detected in green) and a lissamin-labeled human CotI DNA probe (red). (A) The green channel detecting the mouse sequences; (B) The red (human Cot1) and blue (DAPI counterstain) channels. (C,D) The detection of major and minor mouse satellite sequences (biotinylated, detected in green) and of the HCV (lissamin-labeled human Cot1 DNA, red), respectively. The left panels show the green channel; on the right the composite images are shown. Next, a codetection was performed of the HCV using a biotine-labeled alphoid 2 probe (green) and of CENP-C proteins by immunostaining (red signal). (E) The composite image (DAPI counterstain). The red CENP-C signal on the HCV is hidden by the strong green alphoid 2 signal. (F) The red channel only, of the image shown in E. Finally, a slide was sequentially hybridized with a FITC-labeled peptide nucleic acid telomere probe (G, green signals) and a lissamin-labeled human alphoid 2 probe (H, red signal). The poor quality of the metaphase inH results from the combination of the different protocols used for FISH with a peptide nucleic acid probe and a DNA probe. A circle shows the position of the HCV.











