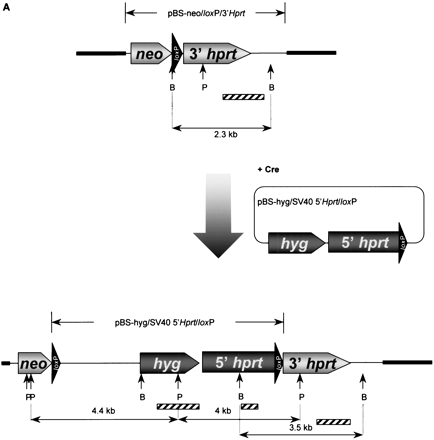
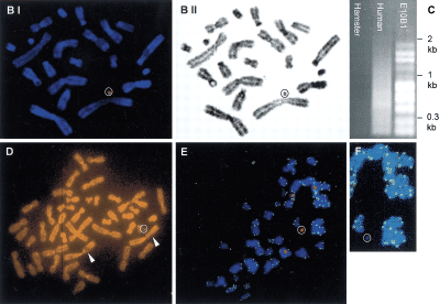
Modification and characterization of the small accessory chromosome (SAC). (A) Structure of the different vectors and strategy for introduction of new sequences into the SAC by Cre-mediated recombination. SAC sequences are indicated with a thick line, vector sequences with a thin line, and loxP sequences with an arrowhead. Neo, neomycine resistance gene driven by a thymidine kinase promoter; hyg , hygromycin resistance cassette driven by the PGK promoter; 5′- and 3′HPRT, human HPRT minigene driven by the SV40 early promoter; P,PstI cleavage site; B, BamHI cleavage site. Fragments used as a probe for Southern blot hybridizations are indicated with a shaded bar (not drawn to scale). (B) FISH with lissamine-labeled human CotI DNA (red signal) and biotine-labeled pBS-Neo/loxP/3′HPRT plasmid (green signal) on a methaphase of hybrid E10B1. The metaphase was counterstained with DAPI. (BI) Pseudocolored image, the circle shows the SAC; (BII) G-like banding derived from the DAPI channel. (C) In the last lane, inter-alu PCR products using E10B1 genomic DNA as a template are shown. Hamster and human genomic DNA were used as a negative and positive control, respectively. (D) FISH with biotin-labeled inter-alu PCR products on a metaphase of the human HT1080 cell line in which the SAC was introduced by MMCT. The circle indicates the SAC; arrowheads show the signals present on chromosome region 1p. (E) FISH with a FITC-(C3TA2)3 peptide nucleic acid probe (green signals) and a lissamine-labeled α-satellite 2 probe (red signals) on metaphase spreads of the SAC+ HT 1080 cell line did not show a green signal on the SAC (circle). (F) Magnification of the SAC shown in E without the red signal of the α-satellite 2 probe.











