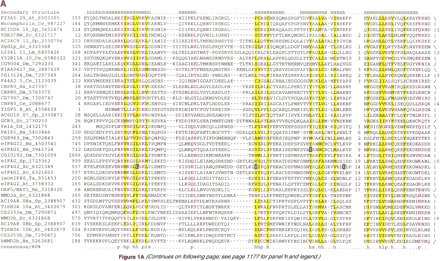
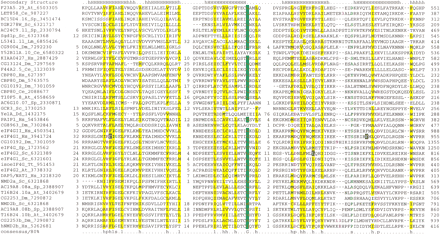

(A) (See pages 1175 and 1176.) Multiple sequence alignment of the predicted NIC domain. (B) Multiple sequence alignment of the predicted MI domain. The proteins are designated by their name, followed by the species abbreviation and GenBank gene identifier. The numbers on either side of the alignment represent the position of the first and last residue of the respective domain in each protein. A consensus secondary structure predicted using thePSIPRED and PHD programs is shown above the alignment. The coloring is based on the 80% consensus in partA and the 85% consensus in part B: h—hydrophobic residues / l—aliphatic residues shaded yellow (YFWLIVMA), c—charged residues colored magenta, p—polar residues colored purple (STQNEDRKH), s—small residues colored green (SAGTVPNHD), t—tiny residues shaded green (GAS), and b—big residues shaded gray (KREQWFYLMI). The point mutations in the human eIF4G1 and yeast eIF4G2 that affect translation are high-lighted in blue in the NIC domain alignment. In the case of the NIC domain alignment 6 distinct families are delineated by brackets to the right of the alignment panel 1. These families are (1) Nucampholin-like, (2) SGD1p-like, (3) KIAA0427-like, (4) CBP80-like, (5) eIF4G-like, and (6) NMD2-like. The species abbreviations are: At— Arabidopsis thaliana, Ce—Caenorhabditis elegans, Dm— Drosophila melanogaster,Mm— Mus musculus, Hs— Homo sapiens, Sc—Saccharomyces cerevisiae, Sp— Schizosaccharomyces pombe, Ta— Triticum aestivum, Pf—Plasmodium falciparum, and Lm—Leishmania major.











