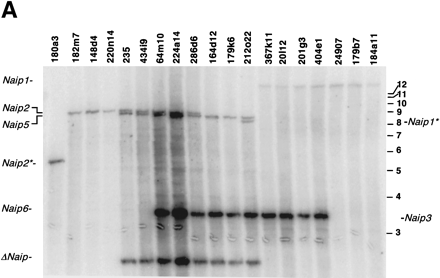
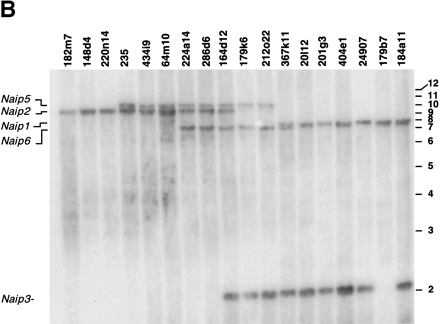
Southern blot analysis of BamHI- and EcoRI-digested BAC DNA identifies six copies of Naip exon 11 and five copies of Naipexon 3. Correlation of the restriction fragments with specificNaip loci was done by comparison of the bands on the BAC blot with predicted fragments from genomic sequence and our previous physical map of the 129 Naip array (data not shown) (Growney et al. 2000). Horizontal bars and numbers indicate position and size (kb) of fragments in 1-kb ladder molecular weight marker (GIBCO). (A) BamHI-digested DNA probed with Naip exon 11 identifies six Naip exon 11 loci. 129 haplotype genomic sequence predicts the observed 14.3-kb fragment mapping toNaip1, a 9-kb fragment mapping to Naip2, an 8.6-kb fragment mapping to Naip5, and a doublet of 3.5 kb and 3.6 kb mapping to Naip6 and Naip3. The remaining 2.2-kb band maps to ΔNaip, as observed in the 129 haplotype (data not shown). Asterisk indicates vector junction fragments mapping toNaip1 and Naip2. (B) EcoRI-digested DNA probed with Naip exon 3 identifies five Naip exon 3 loci. 129 haplotype genomic sequence predicts the observed 10.2-kb fragment mapping to Naip5, an 8.5-kb fragment mapping toNaip2, a 7.5-kb fragment mapping to Naip1, a 7.2-kb fragment mapping to Naip6, and 2.1-kb mapping toNaip3.











