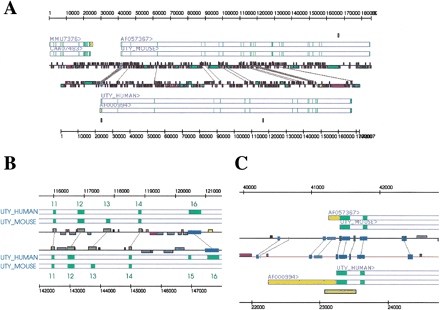
Analysis of the mouse and human UTY genomic regions. (A) Overall view of the region. The mouse sequence (AC006508) is shown at the top and the human sequence (AC006376) at the bottom. Gene structures are the result of analysis with blEst_genome and Blastwise. The sequence IDs of the RNA and protein entries from EMBL, SWISSPROT, and TREMBL that have been aligned to the genomic sequences are shown above the gene structure representations. Regions of similarity detected by BLASTN/MSPcrunch and DBA are connected by gray lines. Regions covered by repeats are represented as colored boxes on the lines representing the sequences. (B) Exons 11–16. Exon positions are derived from GeneWise alignments of the mouse and human protein sequences (Swissprot id: UTY_MOUSE and UTY_HUMAN) with the two genomic sequences as indicated. Dark-yellow blocks connected by gray lines are conserved regions found by BLASTN/MSPcrunch, blue boxes are regions found by DBA. The mouse sequence uses exon 13 but not exons 15 and 16. The human sequence uses exons 15 and 16 but not exon 13. Exon 13 is conserved in both species and parts of exon 16 is also conserved in both species, whereas exon 15 is not. The conservation of these exons indicates the possibility of alternatively spliced transcripts in both species. (C) Conservation of noncoding regions of the 5′ end of the UTY gene. Blocks in different shades of blue connected by gray lines are conserved regions found by DBA. The upstream region contains one conserved block upstream of a possible TATA box (short light-blue region). The 5′ UTR contains two conserved blocks, and the first and second introns contain one conserved region each that are separate from the conservation seen around the exons. These blocks contain several conserved transcription factor-binding sites (see text). A predicted CpG island represented by a yellow box covers parts of the first exon and intron in the human sequence.











