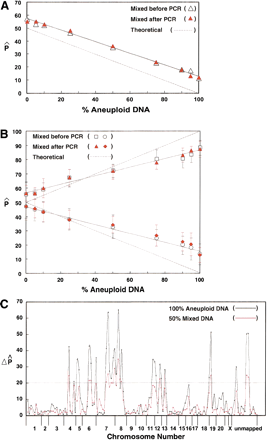
Test of array-based difference detection in heterogeneous populations. The DNA derived from the aneuploid population was mixed into DNA derived from the same patient's normal cells with increasing percentages of 0%, 5%, 10%, 25%, 50%, 75%, 90%, 95%, and 100%. The mixing was performed either prior to or after the locus-specific multiplex PCR and the labeling PCR. (A) The behavior of a single locus with increasing amounts of aneuploid DNA. (Red solid triangles and open black triangles) P̂ Values for the sample mixed before and after PCR, respectively; (solid black line) simple linear fit for the two sets of data. The broken lines are theoretical, indicating what would be expected in an ideal case. (B) The behavior of the average of 13 or 15 loci with increasing amounts of aneuploid DNA. (Open black squares and circles) Average P̂ values from either 13 loci (changed in the AA direction) or 15 loci (changed in the BB direction) for the samples mixed before the PCR; (red solid triangles and diamond) averageP̂ values for the samples mixed after the PCR. The error bars represent the standard deviation of the average P̂values from either 13 or 15 markers. (Solid and broken lines) as described in Fig. 8 A. (C) Comparison of genome-wide difference scans for the 50% mixture and the 100% aneuploid samples. (Pink and black lines) Scans for the 50% mixture and 100% aneuploid samples, respectively. Only informative markers are shown (410 markers passed the quality analysis for all the mixtures and 138 markers were informative for this individual).











