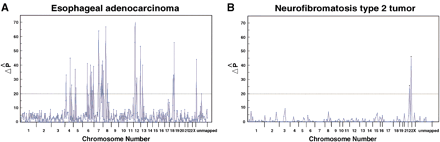
Genome-wide representation of the SNP-based analysis. (A) Genome-wide allelic imbalance detection using SNP markers in the same esophageal adenocarcinoma aneuploid population from the reproducibility experiment (Fig. 4). Of 558 SNP markers 470 passed the quality analysis and 150 of the 470 markers were informative for this individual. Chromosomal regions with SNPs showing a ‖ ΔP̂ ‖ ≥ 20 were independently checked with STRs that lie within the SNP region or flank the SNP loci. The ‖ ΔP̂ ‖ values for all loci including non-informative SNPs are shown. (B) Genome-wide difference detection in NF-2 tumors. For this experiment, an older version of the SNP arrays containing 250 SNPs was used, with 167 of the 250 SNPs passing the quality analysis and 63 of the 167 markers being informative. The distance between tick marks on the x-axis is defined by the number of SNPs on each chromosome (based on the Whitehead Institute SNP map). The values on the y-axis are the difference in ‖ ΔP̂ ‖ values between normal and tumor samples. The dashed line indicates the threshold value (‖ ΔP̂ ‖ ≥ 20), as described in the Data Analysis section.











