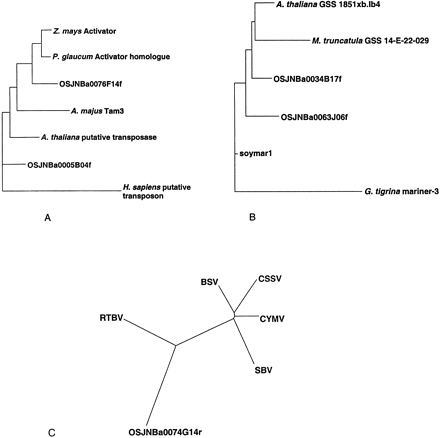
Phylogenies of TE homologs in the rice STC database. All phylogenies were constructed using the Fitch-Margoliash (1967) method. (A) Phylogeny of Activator-like protein sequences, derived from a partial-length multiple sequence alignment of 197 amino acids. Sequences from top to bottom are maizeActivator (P08770), pearl millet Activator homolog (1091678), rice STC OSJNBa0076F14f, snap dragon Tam3 (S13518),Arabidopsis putative transposase (AAD24567), rice STC (OSJNBa0005B04f), and human putative transposon (NP_004720). Translations of rice STCs were obtained from TFASTX alignments of maizeActivator (P08770) with the STC database. (B) Phylogeny of Mariner-like protein sequences, derived from a partial-length multiple sequence alignment of 107 amino acids. Sequences from top to bottom are Arabidopsis thaliana genome survey sequence 1851xb.lb4 (AF005799),Medicago truncatula genome survey sequence 14-E-22–029 from the Crop Biotechnology Center, Texas A & M University (AQ841462), rice STCs OSJNBa0034B17f and OSJNBa0063J06f, soybean Marinerelement soymar1 (AAC28384), and flatworm Girardia tigrina mariner-3 (CAA56859). Translations of rice STCs and other genome survey sequences were obtained from TFASTX alignments ofsoymar1 with the rice STC database and GenBank. (C) Phylogeny of pararetrovirus coat protein sequences, derived from a partial-length multiple sequence alignment of 220 amino acids. Sequences are from rice tungro bacilliform virus (RTBV, AAD30194), banana streak virus (BSV, CAA05264), cacao swollen shoot virus (CSSV, AAA03171), Commelina yellow mottle virus (CYMV, S11479), sugarcane bacilliform virus (SBV, S27938), and rice STC OSJNBa0074G14r. Translation of rice STC was obtained from a TFASTX alignment with CYMV protein 3 (S11479).











