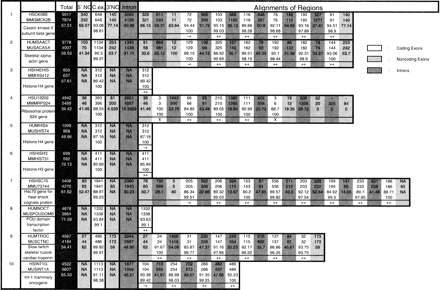
Comparative Analysis of Human and Mouse Loci
|
-
(The complete table is available online as supplemental material at theGenome Research Website: www.genome.org.) In this table we report a structural comparison of 117 orthologous human and mouse genomic loci. We also report the exon prediction performance ofROSETTA on each of these loci. Each entry in the table, numbered 1–117, is a pair of orthologous loci. In the first column, the GenBank LOCUS of the human entry, followed by the GenBank LOCUS of the mouse entry, followed by short descriptions of the genes, are given. The following columns have the following meanings, depending on the rows: (1) first row corresponds to the human entry; (2) second row corresponds to the mouse entry; (3) third row corresponds to nucleotide sequence similarity; (4) fourth row, when applicable, corresponds to amino acid similarity; and (5) fifth row, when applicable, corresponds toROSETTA predictions. Thus the columns have the following meaning: (1) third column, colored dark, corresponds to the total size for human and mouse and the total sequence similarity using the GLASS alignment; (2) fourth column corresponds to the sizes and nucleotide similarity of the 5′-UTRs; (3) fifth column corresponds to the sizes, nucleotide, and protein similarity of the translated regions; (4) sixth column corresponds to the sizes and nucleotide similarity of the 3′-UTRs; (5) seventh column corresponds to the sizes and nucleotide similarity of the introns; and (6) the rest of the columns correspond to the sizes, nucleotide similarity, and protein similarity plus ROSETTA predictions whenever applicable. The color shading the regions indicates the type of the regions: coding exons (white); noncoding exons (light gray); and introns (medium dark gray). The ROSETTA predictions are indicated as follows: (++) coding exon predicted correct on both ends; (+−) coding exon predicted correct only on the 3′-end; (−+) coding exon predicted correct only on the 5′-end. (−−) coding exon was not missed totally, but both 3′- and 5′-boundaries were wrongly predicted; and (X) coding exon was missed altogether. Structurally unusual cases, such as when two coding exons in human correspond to one in mouse, can readily be seen in the table. For instance, entry 30 has such a situation. Coding exons 5 and 6 in human can be seen to correspond to coding exon 5 in mouse.











