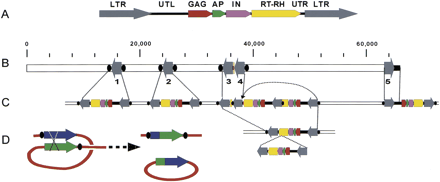
Retrotransposon BARE-1 insertions in the 66-kb contig. (A) Organization of a full-length 8.9-kb BARE-1element as described in the text. The unit is drawn in the sense direction. (B) Organization of the BARE-1 domains on the contig. Arrows represent LTRs numbered as in the text. Black ovals (not to scale) indicate direct repeats found adjacent to the LTRs that were created upon BARE-1 insertion. (C) Proposed ancestral state of current genomic organization. Dotted lines indicate inter-/intrachromosomal recombination events resulting in loss ofBARE-1 internal regions. The LTR-3 and -4 complex resulted from the insertion of one BARE-1 into another followed by recombination between two of the LTRs (shown by the arrow). LTR-5 and part of a UTL terminate the clone. (D) Model for recombination between two LTRs, resulting in a single recombinant LTR in the genome and a closed circle, bearing an LTR and the internal domain, which is then lost.











