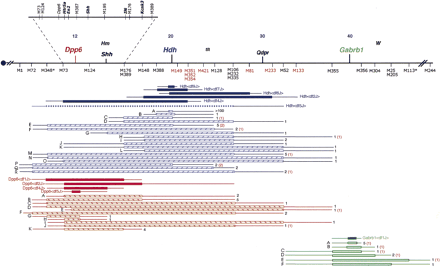
Deletion complexes on chromosome 5. The proximal region of chromosome 5 is depicted as a horizontal line, with the centromere (circle) on the left. Map positions (in centimorgans) as reported in the MGD are indicated, along with the approximate locations of certain landmark loci, such as hammertoe (Hm), Qdpr, and Kit, also known as dominant white spotting (W). Shh is sonic hedgehog homolog, within which a simple sequence repeat was used to type deletions (see Methods). Microsatellite loci, indicated under the map, have been abbreviated by exchanging the prefix “M” for “D5Mit.” The map positions are based on three sources: MGD values, deletion mapping, and RH mapping. As discussed in the text, analysis of deletion breakpoints revealed two disagreements in microsatellite order with MGD; these markers are indicated with asterisks, and in both cases, the MGD order juxtaposes these two markers with the adjacent, proximal microsatellites on this map. A region surroundingDpp6 is expanded to integrate data obtained by high resolution RH mapping, as described in the text. Deletions are indicated as horizontal rectangles, either solid, striped, or unfilled, and are color coded with the locus at which they were induced (red,Dpp6; blue, Hdh; green, Gabrb1). The amount of DNA known to be absent in each deletion is spanned by the rectangles. The thin lines extending from the ends of the rectangles indicate the regions in which the deletion breakpoints reside. Those deletions that have been converted into stocks of mice are represented by solid rectangles, and associated allele names are given (the bracketed text corresponds to superscripting). Deletions existing only in ES cells are represented by the striped or unfilled rectangles. In this latter case, the number of independent ES cell lines containing a particular class of deletion (currently indistinguishable with the markers used in this study) is indicated on the right. In some cases, red numbers in parentheses next to these indicate the number of ES cell lines in this deletion class that failed to produce germline chimeras. An ES cell clone containing the Hdhdf5J deletion, indicated by a dashed line, generated a germline chimera that only transmitted the nondeleted chromosome to its offspring. The relative positions of D5Mit421, tlt (tilted), andD5Mit128 are taken from Ying et al. (Ying et al. 1999). Microsatellite markers that are polymorphic between 129 and B6 are in black, whereas those that are not are in red. The following markers had a 129 allele size smaller than B6: D5Mit1, 25, 52, 72, 73, 106, 124, 148, 176, 232, 335, 348, 356, 388,and Dpp6Rep3. The remaining SSRs had larger 129 alleles.











