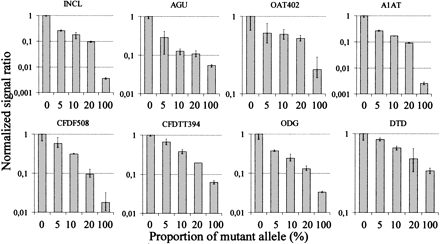
Detection of minority mutations in mixed samples with an excess of normal sequence by dual-color, allele-specific primer extension reactions. A control sample, which is a normal homozygote for all the tested mutations (0% mutant allele), was first analyzed on all the arrays by extension with CY3-labeled dNTPs. The same arrays were then subjected to a second round of extension, in which the mixed samples containing varying amounts of mutant allele were analyzed using CY5-labeled dNTPs in the allele-specific reactions. The acronyms for the mutations above each diagram are as in Figure 3. The proportion of mutant allele (%) in the samples is given below the diagrams. The CY-5 signal ratios between the mutant and normal alleles in the mixed samples normalized for the corresponding CY-3 ratios in the control sample and further normalized to 1 are given on the y-axis. The error bars show range of signal ratios in triplicate.











