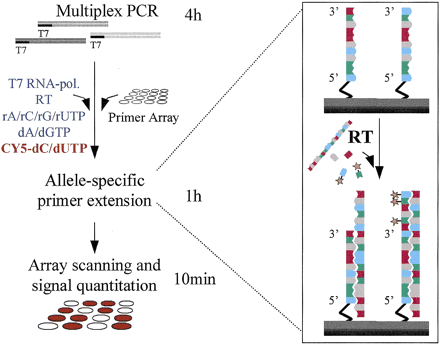
Steps and principle of allele-specific extension on primer arrays. Multiplex PCRs yielding amplicons tailed with 5′-T7 RNA polymerase promoter sequence are performed. The PCR products are added directly to the primer array, along with the reaction mixture, which contains both T7 RNA polymerase and rNTPs to generate RNA targets from the PCR products and RT and labeled dNTPs for the actual allele-specific genotyping reaction, which is illustrated in the inset on the right column. For genotyping a pair of allele-specific detection primers for each mutant or polymorphic site immobilized through their 5′ end are used. The two primers differ at their 3′ end, which is complementary to either of the variant alleles. The RT extends the immobilized detection primers with labeled dNTPs in a template-dependent manner. After the reaction, fluorescence scanning of the arrays and quantitation of the fluorescent signals are carried out using a commercial confocal scanner and software. The fluorescent signals from each primer pair are compared, and the signal ratios fall into distinct categories defining the genotype at each site. The timescale for the reaction steps is illustrated beside the left column of the figure.











