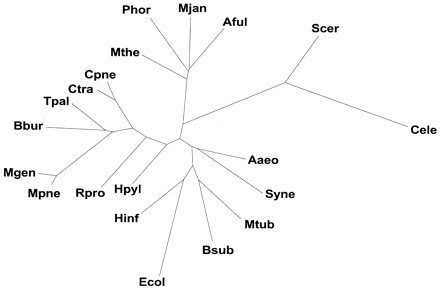
Prospects for the future. The figure shows a 20-genome tree based on the occurrence of folds. This is similar to Figure 3A. The unit in the SCOP classification that was used was the structural superfamily rather than the fold. For eight genome occurrence trees there is no difference between one made at the fold or superfamily level. However, for the 20-genome tree this distinction matters. The additional species names in the 20-genome tree are: Aaeo (Aquifex aeolicus), Aful (Archaeoglobus fulgidus), Bsub (Bacillus subtilis), Bbur (Borrelia burgdorferi), Cpne (Chlamydia pneumoniae), Ctra (Chlamydia trachomatis), Ecol (Escherichia coli), Hinf (Haemophilus influenzae), Hpyl (Helicobacter pylori), Mthe (Methanobacterium thermoautotrophicum), Mjan (Methanococcus jannaschii), Mtub (Mycobacterium tuberculosis), Mgen (Mycoplasma genitalium), Mpne (Mycoplasma pneumoniae), Phor (Pyrococcus horikoshii), Rpro (Rickettsia prowazekii), Scer (Saccharomyces cerevisiae), Syne (Synechocystis sp.), and Tpal (Treponema pallidum).











