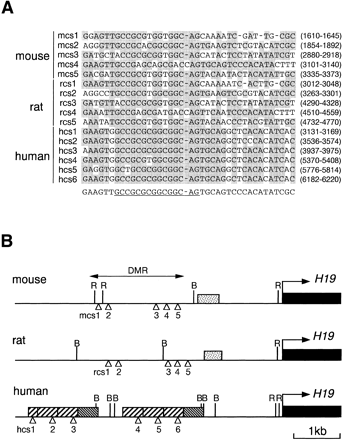
Evolutionarily conserved 39-bp sequence elements in the upstream DMR ofH19. (A) Sequence alignment of the elements conserved in human (hcs1–hcs6), mouse (mcs1–mcs5) and rat (rcs1–rcs5). The bases identical to the consensus sequence (bottom) are shaded. The highly conserved core of the consensus sequence is underlined. Nucleotide positions (in parentheses) follow numbering schemes forAF087017 (human), AF049091 (mouse), and AF043428 (rat). (B) Map of the upstream DMR of human, mouse, and rat H19. (Stippled boxes) G-rich repeat region found in rodents; (hatched boxes) two types of the human 400-bp repeat sequences; (open arrowheads) locations of the conserved 39-bp sequences. B, BamHI; R,EcoRI.











