cDNA Selection Clones, Configs, and Exon Trapping Products
|
-
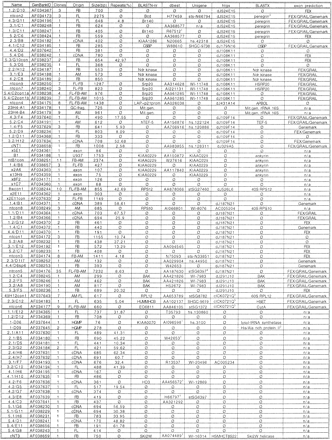
Details of cDNA selection clones, contigs, and exon trapping products ordered from centromere to telomere. Clones below the black bar possibly fall outside the presented contig. (Clones) Number of clones that form a contig. (Origin) FB, fetal brain; FL, fetal liver; AM, adult muscle; cDNA, from cDNA selection, but origin could not be determined; HGMP, from the gridded HGMP library; exon, from the exon trapping; GC, frag., from the GC-rich isolation experiment. (Repeats) Percentage of repeats, identified by RepeatMasker. (BLASTN-nr) BlastN hits in the nonredundant sequence databases. (dbest) BlastN hits in the EST sequence databases. (htgs) BlastN hits in the high throughput sequence databases. (Unigene) The identified ESTs belong to a Unigene contig. (BLASTX) Best match is shown in each case. (Exon prediction) Predictions of the FEX, GRAIL, and Genemark algorithms from NIX.
-
For these matches P > le-100. For R67512,P = 7e-20; AI096598, P = 2e-10; W42653,P = 2e-43; AA456; P > 2e-16; H66797,P = 4e-15; AA074489, P = 3e-54.
-
P = 4.5e-7.
-
P = 2.1e-5.
-
Also WI-15639.
-
Also cICK0921Q.











