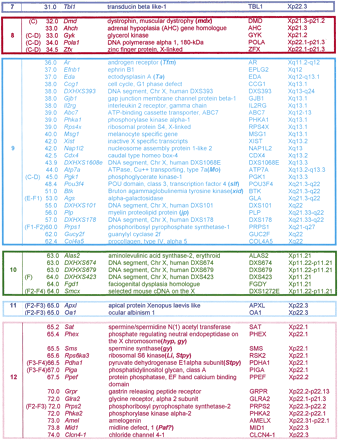
List of X-Linked Genes and Conserved Sequences

|
|
-
Included are only those loci that have been regionally mapped on both the human and mouse X chromosomes with sufficient resolution to contribute to the man–mouse comparative map (see Fig. 1). This list is given in order of loci from the mouse centromere to the telomere. The rectangles encase conserved segments, in which locus order is preserved in man and mouse, with the exception of a small inversion around Xist (see text). Note that the relative order of segments 1–3 has not been established, and eIF2γ (Ehrman et al. 1998) has not been sufficiently well positioned in the mouse to include here, although it might represent a novel conserved segment. The location of each locus on the mouse X chromosome is given in column 1 and indicates the distance in cM from the centromere. When a locus also has been mapped by in situ hybridization, the cytogenetic location on the mouse X chromosome is given in brackets before the genetic map position. Mapping data are taken from the literature (Boyd et al. 1998), except for the Sat locus (H.J. Blair, unpubl.). The cytogenetic position of each locus is given for its location on the human X chromosome as, because of the large number of translocation breakpoints and rearrangements affecting the human X chromosome, this provides a more useful reference than a genetic map position. For a visual appreciation of the relative physical size of the conserved segments and their locations on the human and mouse chromosomes, see Fig. 1.











