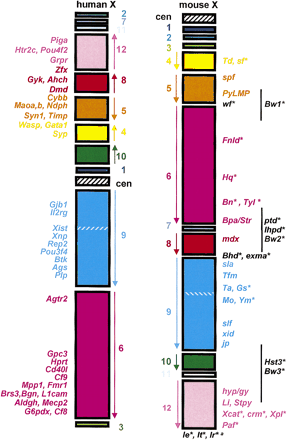
The comparative phenotype map of the human and mouse X chromosomes. Each conserved segment, in which the order of loci is the same in mouse and man, is indicated by a colored rectangle. The loci that define each segment are given in Table 1; note that the order of segments1, 2, and 3 has not been established and is arbitrary. The segments are numbered from 1–12 from the centromere to the telomere on the mouse X chromosome and the order of loci indicated by an arrow alongside each block. The hatched region within segment 9 represents the 600-kb region that is inverted around the Xist locus (see text). The centromeric regions are indicated by black and white hatched rectangles; there is, as yet, no known evidence for evolutionary conservation of centromeric sequences between mouse and man. The Xp pseudoautosomal region, which has a complex evolutionary history (Blaschke and Rappold 1997), is not included because it is outside the scope of this review, as loci in this region do not exhibit X-linked inheritance. Classical mutants carrrying spontaneous and induced mutations and variants (Table 2) are indicated on the mouse X chromosome, and targeted mutations (Table 3) are positioned on the human X chromosome for clarity. Names in black text are (1) known to have an X-linked inheritance pattern but have not been positioned on the X (e.g., Ie, It), or (2) have not been subjected to high-resolution mapping (e.g., Bw1, the black line indicates the probable region in which the locus lies), or (3) have been mapped to evolutionary breakpoint positions and therefore the position of the human homolog cannot be defined (e.g.,exma, Bhd, wf). For those traits marked with an asterisk (*), the gene responsible is not known; for all other traits the underlying lesion has been defined.
a(Ir) Immune response genes that are X linked (see Table 2 footnote).











